User Tools
This is an old revision of the document!
Table of Contents
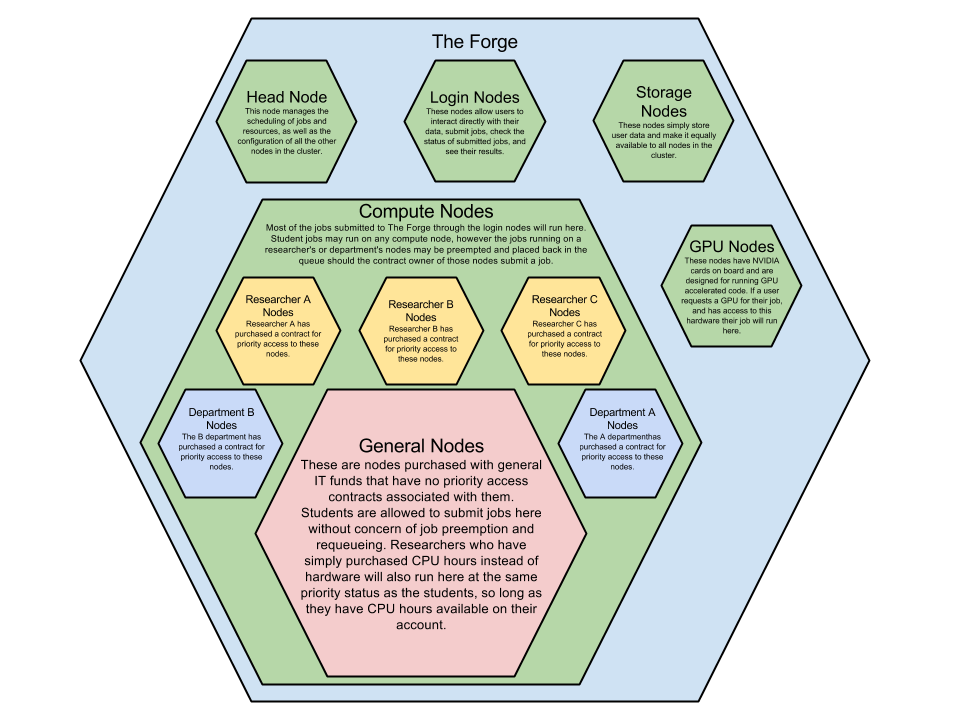
The Forge
System Information
Software
The Forge was built with rocks 6.1.1 which is built on top of CentOS 6.5. With the Forge we made the conversion from using PBS/Maui to using SLURM as our scheduler and resource manager.
Hardware
Management nodes
The head node and login node are Dell commodity servers configured as follows.
Dell R430: Dual 8 core Haswell CPUs with 64GB of DDR4 ram and dual 1TB 7.2K rpm drives in RAID 1
Compute nodes
The newly added compute nodes are all SuperMicro super servers TP-2028 chassis configured as follows.
SuperMicro TP-2028: 4 node chassis with each node containing dual 16 core Haswell CPUs with 256 GB DDR4 ram and 6 300GB 15K SAS drives in raid 0.
Storage
The Forge home directory storage is a Dell NSS high availability storage solution. This storage will provide 480 TB of raw storage, 349 TB of that is available for users, limited to 50GB per user, which can be expanded upon request with proof of need. This volume is not backed up, we do not provide any data recovery guarantee in the event of a storage system failure. System failures where data loss occurs are rare, but they do happen. All this to say, you should not be storing the only copy of your critical data on this system.
Policies
Under no circumstances should your code be running on the login node.
You are allowed to install software in your home directory for your own use. Know that you will *NOT* be given root/sudo access, so if your software requires it you will not be able to use that software. Contact ITRSS about having the software installed cluster-wide.
User data on the Forge is not backed up meaning it is your responsibility to back up important research data to a location off site via any of the methods in the moving_data section.
If you are a student your jobs can run on any compute node in the cluster, even the ones dedicated to researchers, however if the researcher who has priority access to that dedicated node then your job will stop and go back into the queue. You may prevent this preemption by specifying to run on just the general nodes in your job file, please see the documentation on how to submit this request.
If you are a researcher who has purchased priority access for 3 years to an allocation of nodes your jobs will primarily run on those nodes, if your job requires more resources than you have available in your priority queue your job will run in the general queue at the same priority as student jobs. You may add users to your allocation so that your research group may also benefit from the priority access.
If you are a researcher who has purchased an allocation of CPU hours you will run on the general nodes at the same priority as the students. Your job will not run on any dedicated nodes and will not be susceptible to preemption by any other user. Once your job starts it will run until it fails, completes, or runs out of allocated time.
Partitions
The Hardware in the Forge is split up into separate groups, or partitions. Some hardware is in more than one partition, for the most part you won't have to define a partition in a job submission file as we have a plugin that will select the best partition for your user for the job to run in. However there are a few cases that you will want to assign a job to a specific partition. Please see the table below for a list of the limits or default values given to jobs based on the partition. The important thing to note is how long you can request your job to run.
| Partition | Time Limit | Default Memory per CPU |
| short | 4 hours | 1000MB |
| requeue | 7 days | 1000MB |
| free | 14 days | 1000MB |
| any priority partition | 30 days | varies by hardware |
Quick Start
Logging in
SSH (Linux)
Open a terminal and type
ssh username@forge.mst.edu
replacing username with your campus sso username, Enter your sso password
Logging in places you onto the login node. Under no circumstances should you run your code on the login node.
If you are submitting a batch file, then your job will be redirected to a compute node to be computed.
However, if you are attempting use a GUI, ensure that you do not run your session on the login node (Example: username@login-44-0). Use an interactive session to be directed to a compute node to run your software.
sinteractive
For further description of sinteractive, read the section in this documentation titled Interactive Jobs.
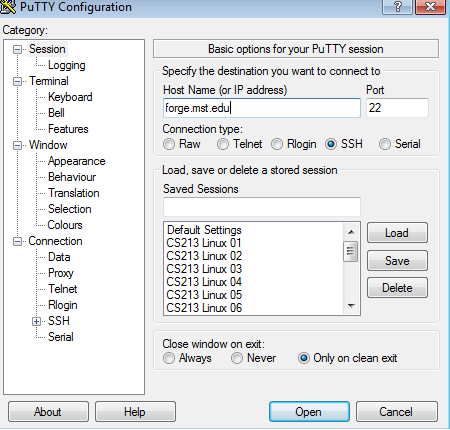
Putty (Windows)
Off Campus Logins
Our off campus logins go through a different host, forge-remote.mst.edu and use public key authentication only, password authentication is disabled for off campus users. To learn how to connect from off campus please see our how to on setting up public key authentication
Submitting a job
Using SLURM is very similar to using PBS/Maui to submit jobs, you need to create a submission script to execute on the backend nodes, then use a command line utility to submit the script to the resource manager. See the file contents of a general submission script complete with comments.
Example Job Script
- batch.sub
#!/bin/bash #SBATCH --job-name=Change_ME #SBATCH --ntasks=1 #SBATCH --time=0-00:10:00 #SBATCH --mail-type=begin,end,fail,requeue #SBATCH --export=all #SBATCH --out=Forge-%j.out # %j will substitute to the job's id #now run your executables just like you would in a shell script, Slurm will set the working directory as the directory the job was submitted from. #e.g. if you submitted from /home/blspcy/softwaretesting your job would run in that directory. #(executables) (options) (parameters) echo "this is a general submission script" echo "I've submitted my first batch job successfully"
Now you need to submit that batch file to the scheduler so that it will run when it is time.
sbatch batch.sub
You will see the output of sbatch after the job submission that will give you the job number, if you would like to monitor the status of your jobs you may do so with the squeue command.
Common SBATCH Directives
| Directive | Valid Values | Description |
| –job-name= | string value no spaces | Sets the job name to something more friendly, useful when examining the queue. |
| –ntasks= | integer value | Sets the requested CPUS for the job |
| –nodes= | integer value | Sets the number of nodes you wish to use, useful if you want all your tasks to land on one node. |
| –time= | D-HH:MM:SS, HH:MM:SS | Sets the allowed run time for the job, accepted formats are listed in the valid values column. |
| –mail-type= | begin,end,fail,requeue | Sets when you would like the scheduler to notify you about a job running. By default no email is sent |
| –mail-user= | email address | Sets the mailto address for this job |
| –export= | ALL,or specific variable names | By default Slurm exports the current environment variables so all loaded modules will be passed to the environment of the job |
| –mem= | integer value | number in MB of memory you would like the job to have access to, each queue has default memory per CPU values set so unless your executable runs out of memory you will likely not need to use this directive. |
| –mem-per-cpu= | integer | Number in MB of memory you want per cpu, default values vary by queue but are typically greater than 1000Mb. |
| –nice= | integer | Allows you to lower a jobs priority if you would like other jobs set to a higher priority in the queue, the higher the nice number the lower the priority. |
| –constraint= | please see sbatch man page for usage | Used only if you want to constrain your job to only run on resources with specific features, please see the next table for a list of valid features to request constraints on. |
| –gres= | name:count | Allows the user to reserve additional resources on the node, specifically for our cluster gpus. e.g. –gres=gpu:2 will reserve 2 gpus on a gpu enabled node |
| -p | partition_name | Not typically used, if not defined jobs get routed to the highest priority partition your user has permission to use. If you were wanting to specifically use a lower priority partition because of higher resource availability you may do so. |
Valid Constraints
| Feature | Description |
| intel | Node has intel CPUs |
| amd | Node has amd CPUs |
| EDR | Node has an EDR (100Gbit/sec) infiniband interconnect |
| FDR | Node has a FDR (56Gbit/sec) infiniband interconnect |
| QDR | Node has a QDR (36Gbit/sec) infiniband interconnect |
| DDR | Node has a DDR (16Gbit/sec) Infiniband interconnect |
| serial | Node has no high speed interconnect |
| gpu | Node has GPU acceleration capabilities |
Monitoring your jobs
squeue -u username JOBID PARTITION NAME USER STATE TIME CPUS NODES 719 requeue Submiss blspcy RUNNING 00:01 1 1
Cancel your job
scancel - Command to cancel a job, user must own the job being cancelled or must be root.
scancel <jobnumber>
Viewing your results
Output from your submission will go into an output file in the submission directory, this will either be slurm-jobnumber.out or whatever you defined in your submission script. In our example script we set this to Forge-jobnumber.out, this file is written asynchronously so it may take a bit after the job is complete for the file to show up if it is a very short job.
Moving Data
Moving data in and out of the forge can be done with a few different tools depending on your operating system and preference.
DFS volumes
Missouri S&T users can mount their web volumes and S Drives with the
mountdfs
command. This will mount your user directories to the login machine under /mnt/dfs/$USER. The data can be copied over with command line tools to your home directory, your data will not be accessible from the compute nodes so do not submit jobs from these directories. Aliases “cds” and “cdwww” have been created to allow you to cd into your s drive and web volume quickly and easily.
Windows
WinSCP
Using winSCP connect to forge.mst.edu using your SSO just as you would with ssh or putty and you will be presented with the contents of your home directory. Now you will be able to drag files into the winscp window and drop them in the folder you want them in and the copying process should begin. It should also work the same in the opposite direction to get data back out.
Filezilla
Using Filezilla you connect to forge.mst.edu using your SSO and you will have the contents of your home directory displayed, drag and drop works with Filezilla as well.
Git
git is installed on the cluster and is recommended to keep track of code changes across your research. See getting started with git for usage guides, Campus offers a hosted private git server at https://git.mst.edu at no additional cost.
Linux
Filezilla
See windows instructions
scp
scp is a command line utility that allows for secure copies from one machine to another through ssh, scp is available on most Linux distributions. If I wanted to copy a file in using scp I would open a terminal on my workstation and issue the following command.
scp /home/blspcy/batch.sub blspcy@forge.mst.edu:/home/blspcy/batch.sub
It will then ask me to authenticate using my campus SSO, then copy the file from my local local location of /home/blspcy/batch.sub to my forge home directory. If you have questions on how to use scp I recommend reading the man page for scp, or check it out online at SCP man page
rsync
rsync is a more powerful command line utility than scp, it has a simpler syntax, and checks to see if the file has actually changed before performing the copy. See the man page for usage details or online documentation
git
See git for windows for instruction, it works the same way.
Modules
An important concept for running on the cluster is modules. Unlike a traditional computer where you can run every program from the command line after installing it, with the cluster we install the programs to a main “repository” so to speak, and then you load only the ones you need as modules. To see which modules are available for you to use you would type “module avail”. Once you find which module you need for your file, you type “module load <module>” where <module> is the module you found you wanted from the module avail list. You can see which modules you already have loaded by typing “module list”.
Here is the output of module avail as of 06/26/2017
------------------------------------------------- /share/apps/modulefiles/mst-mpi-modules ------------------------------------------------- intelmpi openmpi/1.10.6/intel/2013_sp1.3.174 openmpi/2.1.0/intel/2015.2.164 (D) mvapich2/gnu/4.9.2/eth openmpi/1.10.6/intel/2015.2.164 (D) openmpi/gnu/4.9.2/eth mvapich2/gnu/4.9.2/ib (D) openmpi/1.10/gnu/4.9.2 openmpi/gnu/4.9.2/ib (D) mvapich2/intel/11/eth openmpi/1.10/intel/15 openmpi/intel/11/backup mvapich2/intel/11/ib (D) openmpi/2.0.2/gnu/4.9.3 openmpi/intel/11/eth mvapich2/intel/13/eth openmpi/2.0.2/gnu/5.4.0 openmpi/intel/11/ib (D) mvapich2/intel/13/ib (D) openmpi/2.0.2/gnu/6.3.0 openmpi/intel/13/backup mvapich2/intel/15/eth openmpi/2.0.2/gnu/7.1.0 (D) openmpi/intel/13/eth mvapich2/intel/15/ib (D) openmpi/2.0.2/intel/2011_sp1.11.339 openmpi/intel/13/ib (D) mvapich2/pgi/11.4/eth openmpi/2.0.2/intel/2013_sp1.3.174 openmpi/intel/15/eth mvapich2/pgi/11.4/ib (D) openmpi/2.0.2/intel/2015.2.164 (D) openmpi/intel/15/ib (D) openmpi openmpi/2.1.0/gnu/4.9.3 openmpi/pgi/11.4/eth openmpi/1.10.6/gnu/4.9.3 openmpi/2.1.0/gnu/5.4.0 openmpi/pgi/11.4/ib (D) openmpi/1.10.6/gnu/5.4.0 openmpi/2.1.0/gnu/6.3.0 rocks-openmpi openmpi/1.10.6/gnu/6.3.0 openmpi/2.1.0/gnu/7.1.0 (D) rocks-openmpi_ib openmpi/1.10.6/gnu/7.1.0 (D) openmpi/2.1.0/intel/2011_sp1.11.339 openmpi/1.10.6/intel/2011_sp1.11.339 openmpi/2.1.0/intel/2013_sp1.3.174 ------------------------------------------------- /share/apps/modulefiles/mst-lib-modules ------------------------------------------------- arpack glibc/2.14 python/modules/argparse/1.4.0 atlas/gnu/3.10.2 hdf/4/gnu/2.10 python/modules/decorator/4.0.11 atlas/intel/3.10.2 hdf/4/intel/2.10 python/modules/hostlist/1.15 boost/gnu/1.55.0 (D) hdf/4/pgi/2.10 python/modules/kwant/1.0.5 boost/gnu/1.62.0 hdf/5/mvapich2_ib/gnu/1.8.14 python/modules/lasagne/1 boost/intel/1.55.0 (D) hdf/5/mvapich2_ib/intel/1.8.14 python/modules/matplotlib/1.4.2 boost/intel/1.62.0 hdf/5/mvapich2_ib/pgi/1.8.14 python/modules/mpi4py/2.0.1 cgns/openmpi_ib/gnu (D) hdf/5/openmpi_ib/gnu/1.8.14 python/modules/mpmath/0.19 cgns/openmpi_ib/intel hdf/5/openmpi_ib/intel/1.8.14 python/modules/networkx/1.11 fftw/mvapich2_ib/gnu4.9.2/2.1.5 hdf/5/openmpi_ib/pgi/1.8.14 python/modules/nolearn/0.6 fftw/mvapich2_ib/gnu4.9.2/3.3.4 (D) mkl/2011_sp1.11.339 python/modules/numpy/1.10.0 fftw/mvapich2_ib/intel2015.2.164/2.1.5 mkl/2013_sp1.3.174 python/modules/obspy/1.0.4 fftw/mvapich2_ib/intel2015.2.164/3.3.4 (D) mkl/2015.2.164 (D) python/modules/opencv/3.1.0 fftw/mvapich2_ib/pgi11.4/2.1.5 netcdf/mvapich2_ib/gnu/3.6.2 python/modules/pylab/0.1.4 fftw/mvapich2_ib/pgi11.4/3.3.4 (D) netcdf/mvapich2_ib/gnu/4.3.2 (D) python/modules/scipy/0.16.0 fftw/openmpi_ib/gnu4.9.2/2.1.5 netcdf/mvapich2_ib/intel/3.6.2 python/modules/sympy/1.0 fftw/openmpi_ib/gnu4.9.2/3.3.4 (D) netcdf/mvapich2_ib/intel/4.3.2 (D) python/modules/theano/0.8.0 lines 1-79 ------------------------------------------------- /share/apps/modulefiles/mst-app-modules ------------------------------------------------- AnsysEM/17/17 cp2k maple/2015 (D) qe CST/2014 (D) cplot maple/2016 qmcpack/3.0 CST/2015 cpmd matlab/2012 qmcpack/2014 CST/2016 dirac matlab/2013a qmcpack/2016 (D) CST/2017 dock/6.6eth matlab/2014a (D) rum OpenFoam/2.3.1 dock/6.6ib (D) matlab/2016a samtools OpenFoam/2.4.x (D) espresso matlab/2017a scipy ParaView fun3d/custom (D) mcnp siesta R fun3d/12.2 mctdh spark SAS/9.4 fun3d/12.3 metis/5.1.0 sparta SU2/3.2.8 fun3d/12.4 molpro/2010.1.nightly spin SU2/4.1.3 (D) fun3d/2014-12-16 molpro/2010.1.25 starccm/6.04 abaqus/6.11-2 gamess molpro/2012.1.nightly (D) starccm/7.04 abaqus/6.12-3 (D) gaussian molpro/2012.1.9 starccm/8.04 (D) abaqus/6.14-1 gaussian-09-e.01 molpro/2015.1.source.block starccm/10.04 abaqus/2016 gdal molpro/2015.1.source.s starccm/10.06 accelrys/material_studio/6.1 geos molpro/2015.1.source starccm/11.02 amber/12 gmp molpro/2015.1 starccm/11.04 amber/13 (D) gnu-tools moose/3.6.4 tecplot/2013 ansys/14.0 gnuplot mpb-meep tecplot/2014 (D) ansys/15.0 (D) gridgen mpc tecplot/2016.old ansys/17.0 gromacs msc/adams/2015 tecplot/2016 ansys/18.0 hadoop/1.1.1 msc/nastran/2014 trinity apbs hadoop/1.2.1 msc/patran/2014 valgrind/3.12 bowtie hadoop/2.6.0 (D) namd/2.9 vasp/4.6 casino lammps/9Dec14 namd/2.10 (D) vasp/5.4.1 (D) cmg lammps/17Nov16 namd/2.12b visit/2.8.2 comsol/4.3a lammps/30July16 (D) octave visit/2.9.0 (D) comsol/4.4 (D) linpack overture vmd comsol/5.2a_deng lsdyna/8.0 packmol vulcan comsol/5.2a_park lsdyna/8.1.hybrid parmetis/4.0.3 comsol/5.2a_yang lsdyna/8.1 (D) proj comsol/5.2a_zaeem maple/16 psi4 ---------------------------------------------- /share/apps/modulefiles/mst-compiler-modules ----------------------------------------------- cilk/5.4.6 gnu/4.9.3 gpi2/1.0.1 intel/2015.2.164 (D) python/2.7.9 cmake/3.2.1 (D) gnu/5.4.0 gpi2/1.1.1 (D) mono/3.12.0 python/3.5.2_GCC cmake/3.6.2 gnu/6.3.0 intel/2011_sp1.11.339 pgi/11.4 python/3.5.2 (D) gnu/4.9.2 (D) gnu/7.1.0 intel/2013_sp1.3.174 pgi/RCS/11.4,v Where: L: Module is loaded D: Default Module Use "module spider" to find all possible modules. Use "module keyword key1 key2 ..." to search for all possible modules matching any of the "keys".
Compiling Code
There are several compilers available through modules, to see a full list of modules run
module avail
the naming scheme for the compiler modules are as follows.
MPI_PROTOCOL/COMPILER/COMPILER_VERSION/INTERFACE e.g openmpi/intel/15/ib is the 2015 intel compiler built with openmpi libraries and is set to communicate over the high speed infiniband interface.
After you have decided which compiler you want to use you need to load it.
module load openmpi/intel/15/ib
Then compile your code, use mpicc for c code and mpif90 for fortran code. Here is an MPI hello world C code.
- helloworld.c
/* C Example */ #include <stdio.h> #include <mpi.h> int main (argc, argv) int argc; char *argv[]; { int rank, size; MPI_Init (&argc, &argv); /* starts MPI */ MPI_Comm_rank (MPI_COMM_WORLD, &rank); /* get current process id */ MPI_Comm_size (MPI_COMM_WORLD, &size); /* get number of processes */ printf( "Hello world from process %d of %d\n", rank, size ); MPI_Finalize(); return 0; }
Use mpicc to compile it.
mpicc ./helloworld.c
Now you should see a a.out executable in your current working directory, this is your mpi compiled code that we will run when we submit it as a job.
Submitting an MPI job
You need to be sure that you have the same module loaded in your job environment as you did when you compiled the code to ensure that the compiled executables will run correctly, you may either load them before submitting a job and use the directive
#SBATCH --export=all
in your submission script, or load the module prior to running your executable in your submission script. Please see the sample submission script below for an mpi job.
- helloworld.sub
#!/bin/bash #SBATCH -J MPI_HELLO #SBATCH --ntasks=8 #SBATCH --export=all #SBATCH --out=Forge-%j.out #SBATCH --time=0-00:10:00 #SBATCH --mail-type=begin,end,fail,requeue module load openmpi/intel/15/ib mpirun ./a.out
Now we need to submit that file to the scheduler to be put into the queue.
sbatch helloworld.sub
You should see the scheduler report back what job number your job was assigned just as before, and you should shortly see an output file in the directory you submitted your job from.
Interactive jobs
Some things can't be run with a batch script because they require user input, or you need to compile some large code and are worried about bogging down the login node. To start an interactive job simply use the
sinteractive
command and your terminal will now be running on one of the compute nodes. The hostname command can help you confirm you are no longer running on a login node. Now you may run your executable by hand without worrying about impacting other users. The sinteractive script by default will allot a 1 cpu for 1 hour, you may request more by using SBATCH directives, e.g.
sinteractive --time=02:00:00 --cpus-per-task=2
will start a job with 2 CPUs on one node for 2 hours.
If you will need a GUI Window for whatever you are running inside the interactive job you will need to connect to The Forge with X forwarding enabled. For Linux this is simply adding the -X switch to the ssh command.
ssh forge.mst.edu -X
For Windows there are a couple X server software's available for use, x-ming and x-win32 that can be configured with putty. Here is a simple guide for configuring putty to use xming.
Job Arrays
If you have a large number of jobs you need to start I recommend becoming familiar with using job arrays, basically it allows you to submit one job file to start up to 10000 jobs at once.
One of the ways you can vary the input of the job array from task to task is to set a variable based on which array id the job is and then use that value to read the matching line of a file. For instance the following line when put into a script will set the variable PARAMETERS to the matching line of the file data.dat in the submission directory.
PARAMETERS=$(awk -v line=${SLURM_ARRAY_TASK_ID} '{if (NR == line) { print $0; };}' ./data.dat)
You can then use this variable in your execution line to do whatever you would like to do, you just have to have the appropriate data in the data.dat file on the appropriate lines for the array you are submitting. See the sample data.dat file below.
- data.dat
"I am line number 1" "I am line number 2" "I am line number 3" "I am line number 4"
you can then submit your job as an array by using the –array directive, either in the job file or as an argument at submission time, see the example below.
- array_test.sub
#!/bin/bash #SBATCH -J Array_test #SBATCH --ntasks=1 #SBATCH --out=Forge-%j.out #SBATCH --time=0-00:10:00 #SBATCH --mail-type=begin,end,fail,requeue PARAMETERS=$(awk -v line=${SLURM_ARRAY_TASK_ID} '{if (NR == line) { print $0; };}' ./data.dat) echo $PARAMETERS
I prefer to use the array as an argument at submission time so I don't have to touch my submission file again, just the data.dat file that it reads from.
sbatch --array=1-2,4 array_test.sub
Will execute lines 1,2, and 4 of data.dat which echo out what line number they are from my data.dat file.
You may also add this as a directive in your submission file and submit without any switches as normal. Adding the following line to the header of the submission file above will accomplish the same thing as supplying the array values at submission time.
#SBATCH --array=1-2,4
Then you may submit it as normal
sbatch array_test.sub
Converting From Torque
With the new cluster we have changed resource managers from PBS/Maui to Slurm, We have provided a script to convert your submission files from PBS directives to slurm. This script, pbs2slurm.py, should be in your executable path and can be used as follows.
pbs2slurm.py < batch.qsub > batch.sbatch
Where batch.qsub is the name of your old submission script file and batch.sbatch is the name of your new submission script file. Please see the table below for commonly used command conversions.
| PBS/Maui Command | Slurm Equivalent | Purpose |
| qsub | sbatch | Submit a job for execution |
| qdel | scancel | Cancel a job |
| showq | squeue | check the status of jobs |
| qstat | squeue | check the status of jobs |
| checkjob | scontrol show job | check the detailed information of a specific job |
| checknode | scontrol show node | check the detailed information of a specific node |
Checking your account usage
If you have purchased a number of CPU hours from us you may check on how many hours you have used by issuing the
usereport
command from a login node, this will show your account's CPU hour limit and the total amount used.
Note this is usage for your account, not your user.
Applications
The applications portion of this wiki is currently a Work in progress, not all applications are currently here, but they will eventually be.
Abaqus
- Default Vesion = 6.12.3
- Other versions available: 6.11.2, 6.14.1, 2016, 2018
This example is for 6.12
The SBATCH commands are explained in the Forge Documentation.
You will need a *.inp file from where you built the model on another system. This .inp file needs to be in the same folder as the jobfile you will create.
Example jobfile:
- abaqus.sub
#!/bin/bash #SBATCH --job-name=sbatchfilename.sbatch #SBATCH --nodes=1 #SBATCH --ntasks=2 #SBATCH --mem=4000 #SBATCH --time=00:60:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=joeminer@mst.edu\ input=inputfile.inp /share/apps/ABAQUS/Commands/abq6123 job=$input analysis cpus=2 interactive
- Replace the sbatchfilename.sbatch with the name of your jobfile.
- Replace the email address with your own.
- Replace the inputfile.inp with the *.inp file that you want to analyze.
- Keep ntasks and cpu equal to the same amount of processors
This job will run on 1 node with 2 processors and allocate 4GB of memory per processor. These node/processor values can change, but be aware that the Abaqus licensing is limited and the job may queue for hours or days before licenses are available, depending on the quantities you selected.
Simple breakdown of Abaqus Licensing:
| Parallel License | Number of Processors used | Tokens Allocated |
|---|---|---|
| 1 | 4 | 20 |
| 2 | 8 | 40 |
| 3 | 16 | 60 |
| 4 | 32 | 80 |
| 5 | 64 | 100 |
Abaqus 2016
module load abaqus/2016
- abaqus2016.sbatch
#!/bin/bash #SBATCH --job-name=sbatchfilename.sbatch #SBATCH --nodes=1 #SBATCH --ntasks=20 #SBATCH --mem=4000 #SBATCH --partition=requeue #SBATCH --time=10:00:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=joeminer@mst.edu unset SLURM_GTIDS input=inputfile.inp time abq2016hf3 job=$input analysis standard_parallel=all cpus=20 interactive
unset SLURM_GTIDS
This is a new variable that must be in place for 2016 to run with MPI.
Chain loading jobs for Abaqus 2016.
- abaqus2016chain.sbatch
#!/bin/bash #SBATCH --job-name=sbatchfilename.sbatch #SBATCH --nodes=1 #SBATCH --ntasks=20 #SBATCH --mem=4000 #SBATCH --partition=requeue #SBATCH --time=10:00:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=joeminer@mst.edu unset SLURM_GTIDS input=inputfile.inp oldjob=oldjobfile.inp abq2016hf3 job=$input oldjob=$oldjob cpus=20 double interacive
Ansys
- Default Version = 15.0
- Other versions available
- 14.0
Running the Workbench
Be sure you are connected to the Forge with X forwarding enabled, and running inside an interactive job using command
sinteractive
before you attempt to launch the work bench. Running sinteractive without any switches will give you 1 cpu for 1 hour, if you need more time or resources you may request it. See Interactive Jobs for more information.
Once inside an interactive job you need to load the ansys module.
module load ansys
Now you may run the workbench.
runwb2
Job Submission Information
Fluent is the primary tool in the Ansys suite of software used on the Forge.
Most of the fluent simulation creation process is done on your Windows or Linux workstation.
The 'Solving' portion of a simulation is where the Forge is utilized.
Fluent will output a lengthy file, based on the simulation being run and that lengthy output file would be used on your Windows or Linux Workstation to do the final review and analysis of your simulation.
The basic steps
1. Create your geometry
2. Setup your mesh
3. Setup your solving method
4. Use the .cas and .dat files, generated from the first three steps, to construct your jobfile
5. Copy those files to the Forge, to your home folder
6. Create your jobfile using the slurm tools on the Forge Documentation page
7. Load the Ansys module
8. Submit your newly created jobfile with sbatch
Serial Example.
I used the Turbulent Flow example from Cornell's SimCafe.
On the Forge, I have this directory structure for this example. Please create your own structure that makes sense to you.
TurbulentFlow/ |-- flntgz-48243.cas |-- flntgz-48243.dat |-- output.dat |-- slurm-8731.out |-- TurbulentFlow_command.txt |-- TurbulentFlow.sbatch
The .cas file is the CASE file that contains the parameters define by you when creating the model.
The .dat file is the data result file used when running the simulation.
The .txt file, is the actual, command equivalent, of your model, in a form that the Forge understands.
The .sbatch file, is the slurm job file that you will use to submit your model for analysis.
The .out file is the output from the run.
The .dat file is the binary (ansys specific) file created during the solution, that could be imported into Ansys back on the Windows/Linux workstation for further analsys.
Jobfile Example.
- TurbulentFlow.sbatch
#!/bin/bash #SBATCH --job-name=TurbulentFlow.sbatch #SBATCH --ntasks=1 #SBATCH --nodes=1 #SBATCH --time=01:00:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=rlhaffer@mst.edu #SBATCH -o Forge-%j.out fluent 2ddp -g < /home/rlhaffer/unittests/ANSYS/TurbulentFlow/TurbulentFlow_command.txt
The SBATCH commands are explained in the Forge Documentation.
The job-name is a name given to help you determine which job is which.
This job will be in the — partition=requeue queue.
It will use 1 node — nodes=1.
It will use 4 processors in one node — ntasks=4.
It has a wall clock time of 1 hour — time=01:00:00.
It will email the user when it begins, ends, or if it fails. –mail-type & –mail-user
fluent is the command we are going to run.
2ddp is the mode we want fluent to use
Modes
The [mode] option must be supplied and is one of the following:
* 2d runs the two-dimensional, single-precision solver
* 3d runs the three-dimensional, single-precision solver
* 2ddp runs the two-dimensional, double-precision solver
* 3ddp runs the three-dimensional, double-precision solver
-g turns off the GUI
Path to the command file we are calling in fluent. < /home/rlhaffer/unittests/ANSYS/TurbulentFlow/TurbulentFlow_command.txt
Contents of command file
This file can get long. As it contains the .cas file & .dat file information as well as saving frequency and iteration count
NOTE, this is all in one line when creating the command file
/file/rcd /home/rlhaffer/unittests/ANSYS/TurbulentFlow/flntgz-48243.cas /file/autosave/data-frequency 20000 /solve/iterate 150000 /file/wd /home/rlhaffer/unittests/ANSYS/TurbulentFlow/output.dat /exit
When the simulation is finished, you will have a Forge-#####.out file that looks something like this:
/share/apps/ansys_inc/v150/fluent/fluent15.0.7/bin/fluent -r15.0.7 2ddp -g
/share/apps/ansys_inc/v150/fluent/fluent15.0.7/cortex/lnamd64/cortex.15.0.7 -f fluent -g (fluent "2ddp -alnamd64 -r15.0.7 -path/share/apps/ansys_inc/v150/fluent")
Loading "/share/apps/ansys_inc/v150/fluent/fluent15.0.7/lib/fluent.dmp.114-64"
Done.
/share/apps/ansys_inc/v150/fluent/fluent15.0.7/bin/fluent -r15.0.7 2ddp -alnamd64 -path/share/apps/ansys_inc/v150/fluent -cx edrcompute-43-17.local:56955:53521
Starting /share/apps/ansys_inc/v150/fluent/fluent15.0.7/lnamd64/2ddp/fluent.15.0.7 -cx edrcompute-43-17.local:56955:53521
Welcome to ANSYS Fluent 15.0.7
Copyright 2014 ANSYS, Inc.. All Rights Reserved.
Unauthorized use, distribution or duplication is prohibited.
This product is subject to U.S. laws governing export and re-export.
For full Legal Notice, see documentation.
Build Time: Apr 29 2014 13:56:31 EDT Build Id: 10581
Loading "/share/apps/ansys_inc/v150/fluent/fluent15.0.7/lib/flprim.dmp.1119-64"
Done.
--------------------------------------------------------------
This is an academic version of ANSYS FLUENT. Usage of this product
license is limited to the terms and conditions specified in your ANSYS
license form, additional terms section.
--------------------------------------------------------------
Cleanup script file is /home/rlhaffer/unittests/ANSYS/TurbulentFlow/cleanup-fluent-edrcompute-43-17.local-17945.sh
>
Reading "/home/rlhaffer/unittests/ANSYS/TurbulentFlow/flntgz-48243.cas"...
3000 quadrilateral cells, zone 2, binary.
5870 2D interior faces, zone 1, binary.
30 2D velocity-inlet faces, zone 5, binary.
30 2D pressure-outlet faces, zone 6, binary.
100 2D wall faces, zone 7, binary.
100 2D axis faces, zone 8, binary.
3131 nodes, binary.
3131 node flags, binary.
Building...
mesh
materials,
interface,
domains,
mixture
zones,
pipewall
outlet
inlet
interior-surface_body
centerline
surface_body
Done.
Reading "/home/rlhaffer/unittests/ANSYS/TurbulentFlow/flntgz-48243.dat"...
Done.
iter continuity x-velocity y-velocity k epsilon time/iter
! 389 solution is converged
389 9.7717e-07 1.0711e-07 2.9115e-10 5.2917e-08 3.4788e-07 0:00:00 150000
! 390 solution is converged
390 9.5016e-07 1.0389e-07 2.8273e-10 5.1020e-08 3.3551e-07 1:11:14 149999
Writing "/home/rlhaffer/unittests/ANSYS/TurbulentFlow/output.dat"...
Done.
Parallel Example
To use fluent in parallel please you need set the PBS_NODEFILE envrionment variable inside your job. Please see example submission file below.
- TurbulentFlow.sbatch
#!/bin/bash #SBATCH --job-name=TurbulentFlow.sbatch #SBATCH --ntasks=32 #SBATCH --time=01:00:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=rlhaffer@mst.edu #SBATCH -o Forge-%j.out #generate a node file export PBS_NODEFILE=`generate_nodefile` #run fluent in parallel. fluent 2ddp -g -t32 -pinfiniband -cnf=$PBS_NODEFILE -ssh < /home/rlhaffer/unittests/ANSYS/TurbulentFlow/TurbulentFlow_command.txt
Interactive Fluent
If you would like to run the full GUI you may do so inside an interactive job, make sure you've connected to The Forge with X Forwarding enabled. Start the job with.
sinteractive
This will give you 1 processor for 1 hour, to request more processors or more time please see the documentation at Interactive Jobs.
Once inside the interactive job you will need to load the ansys module.
module load ansys
Then you may start fluent from the command line.
fluent 2ddp
will start the 2d, double precision version of fluent. If you've requested more than one processor you need to first run
export PBS_NODEFILE=`generate_nodefile`
Then you need to add some switches to fluent to get it to use those processors.
fluent 2ddp -t## -pethernet -cnf=$PBS_NODEFILE -ssh
You need to replace the ## with the number of processors requested.
Ansys Mechanical
Tested for version 18.1
- round.sbatch
#!/bin/bash #SBATCH --job-name=round #SBATCH --ntasks=10 #SBATCH --mem=20000 #SBATCH -p=requeue #SBATCH --time=00:60:00 --comment=#SBATCH --mail-type=begin --comment=#SBATCH --mail-type=end #SBATCH --export=all #SBATCH --out=Forge-%j.out module load ansys/18.1 time ansys181 -j round -b -dis -np 10 < /home/rlhaffer/unittests/ANSYS/roundthing/round.log
ansys mechanical log file
- round.inp
/INPUT,'round','inp',,0,0 /SOLU SOLVE FINISH
AnsysEM
- Default Version = 17.0
- Also Called Ansys Electronics Desktop
- ansysem.sub
#!/bin/bash #SBATCH --job-name=jobfile.sbatch #SBATCH --partition=requeue #SBATCH --ntasks=1 #SBATCH --cpus-per-task=20 #SBATCH --time=01:00:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=joeminer@mst.edu module load AnsysEM/17 time ansysedt -ng -batchsolve Model_Cluster_Test_v2.aedt
CMG
Computer Modeling Group - This is a WORKING SECTION - Edits are actively taking place.
Default version = 2015
Constructing the jobfile
From the Windows installation the .dat model will be needed.
Please follow the general Forge documentation for submission techniques.
After logged into The Forge, run:
module load cmg
This loads environment variables needed to for CMG to function.
Copy your .dat file from your Windows system to your cluster home folder. This is explained in the Forge documentation.
Example sbatch jobfile
- cmg.sub
#!/bin/bash #SBATCH --job-name=batchfilename.sbatch #SBATCH --nodes=1 #SBATCH --ntasks=7 #SBATCH --mem=17000 #SBATCH --time=10:00:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=joeminer@mst.edu #SBATCH --export=ALL RunSim.sh gem 2015.10 GasModel4P.dat -parasol 7
| RunSim.h | the CMG sim script that allows the simulation to run on multiple processors |
| gem | the solver specified |
| 2015.10 | version of the software used |
| GasModel4P.dat | the model to be analyzed |
| -parasol 7 | this is the command switch to tell CMG there are processors to use. In this example, 7 processors were allocated. |
CMG licensing
When running the above example, we discovered how CMG’s licensing really works.
We have varying quantities of licenses for different features and each seat of those licenses equates to 20 solver tokens that will be allocated to jobs on the cluster.
For this example..
20 tokens of the solve-stars solver are being used, because we have an instance of the solver open.
35 tokens of the solve-parallel solver are being used, because we have asked for 7 processors.
We currently have 8 seats for Stars and 8 seats for Parallel. This equates to 160 tokens of each. Out of that 160 tokens, your job can take at least 20 tokens from the stars solver, and up to 160 tokens of the parallel solver.
160 tokens of the parallel solver actually equates to 32 processors when building the job file.
So, the max processors requested is 32.
Token Scenario example
If a student submits a parallel job asking for 32 processors (all 160 tokens) and all 160 are free, the job will begin running.
If a second student then submits a job asking for 10 processors (50 tokens) but all 160 are currently in use, the job will fail. In the output file of the failed job, it will indicate that the job failed because it did not have enough licenses to run.
If this happens, the best suggestion is to modify your jobfile with a smaller number of processors. This may take a few iterations until you find a number of processors that will work.
There is no efficient way to know how many licenses are in use.
Licensing for CMG is limited. Some features have 2160 tokens, some only 100.
Please be patient when submitting jobs as they could fail in this no license manner.
The jobs tested only indicate the solve-star and solve-parallel are being used. We have not seen a test case using these other features:
gem_forgas gem_gap imex_forgas imex_gap imex_iam stars_forgas stars_gap winprop builder cmost_studio dynagrid solve_university rlm_roam
CST
- (Computer Simulation Techonolgy)
- Default version = 2014
- Other versions available = 2015
Job Submission information
CST is an electromagnetic simulation solver, primarily used by the EMCLab.
Steps.
- Build your simulation on one of the Windows workstations availble to you.
- Copy your .cst file to your home directory on the Forge.
- Write you job submission file.
- Either use the default CST version of 2014 or use 2015 by loading its module file.
module load CST/2015
Example Job File
- CST.sub
#!/bin/bash #SBATCH --job-name=CST_Test #SBATCH --nodes=1 #SBATCH --ntasks=2 #SBATCH --mem=4000 #SBATCH --partition=cuda #SBATCH --gres=gpu:2 #SBATCH --time=08:00:00 #SBATCH --mail-type=FAIL #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-user=joeminer@mst.edu cst_design_environment /home/userID/path/to/.cst/file/example.cst --r --withgpu=2
This job will use 1 node, 2 processors from that node, 4GB of memory, the cuda queue, 2 gpus, 8 hours of wall clock time and email the user when the job starts, ends, or fails.
This will output the results in the standard CST folder structure:
/path/to/.cst/file/example |-- DC |-- Model |-- ModelCache |-- Model.lok |-- Result |-- SP `-- Temp
Cuda
If you would like to use CUDA programs or GPU accelerated code you will need to use our GPU nodes. Access to our GPU nodes is granted on a case by case basis, please request access to them through the help desk or submit a ticket at http://help.mst.edu.
Our login nodes don't have the CUDA toolkit installed so to compile your code you will need to start an interactive job on these nodes to do your compilation.
sinteractive -p cuda --time=01:00:00 --gres=gpu:1
This interactive session will start on a cuda node and give you access to one of the GPUs on the node, once started you may compile your code and do whatever testing you need to do inside this interactive session.
To submit a job for batch processing please see this example submission file below.
- cuda.sub
#!/bin/bash #SBATCH -J Cuda_Job #SBATCH -p cuda #SBATCH -o Forge-%j.out #SBATCH --nodes=1 #SBATCH --ntasks=1 #SBATCH --gres=gpu:1 #SBATCH --time=01:00:00 ./a.out
This file requests 1 cpu and 1 gpu on 1 node for 1 hour, to request more cpus or more gpus you will need to modify the values related to ntasks and gres=gpu. It is recommended that you at least have 1 cpu for each gpu you intend to use, we currently only have 2 gpus available per node. Once we incorporate the remainder of the GPU nodes we will have 7 gpus available in one chassis.
Espresso
- Espresso is an integrated suite of Open-Source computer codes for electronic-structure calculations and materials modeling at the nanoscale. It is based on density-functional theory, plane waves, and pseudopotentials.
- Default version - 5.2.1
All of the available pseudopotential files .UPF as of 12/14/2015, are copied into the pseudo install folder.
Before running an Espresso job, run:
module load espresso
Example:
The sbatch file is the job file you will run on the Forge HPC.
The .pw.in is the input file you create, that you will reference in the jobfile.
In the job file, change the path to your input file.
In the .pw.in file, change the paths of outdir, wfcdir to a location in your Forge home directory and change the pseudo_dir path as in the example below.
- FeGeom2BU.sbatch
#!/bin/bash #SBATCH --job-name=FeGeomBU #SBATCH --ntasks=20 #SBATCH --time=0-04:00:00 #SBATCH --mail-type=begin,end,fail,requeue #SBATCH --out=forge-%j.out mpirun pw.x -procs 20 -i /path/to/your/inputfile/FeGeom2BU.pw.in
- FeGeom2BU.pw.in
&CONTROL title = 'Na3FePO4CO3 geom+U' , calculation = 'vc-relax' , outdir = '/path/to/your/inputfile' , wfcdir = '/path/to/your/inputfile' , pseudo_dir = '/share/apps/espresso/espresso-5.2.1/pseudo' , prefix = 'Na3FePO4CO3Geom' , verbosity = 'high' , / &SYSTEM ibrav = 12, A = 8.997 , B = 5.163 , C = 6.741 , cosAB = -0.0027925231 , nat = 11, ntyp = 5, ecutwfc = 40 , ecutrho = 400 , lda_plus_u = .true. , lda_plus_u_kind = 0 , Hubbard_U(1) = 6.39, space_group = 11 , uniqueb = .false. , / &ELECTRONS conv_thr = 1d-6 , startingpot = 'file' , startingwfc = 'atomic' , / &IONS ion_dynamics = 'bfgs' , / &CELL cell_dynamics = 'bfgs' , cell_dofree = 'all' , / ATOMIC_SPECIES Fe 55.93300 Fe.pbe-sp-van_mit.UPF P 30.97400 P.pbe-van_ak.UPF C 12.01100 C.pbe-van_ak.UPF O 15.99900 O.pbe-van_ak.UPF Na 22.99000 Na.pbe-sp-van_ak.UPF ATOMIC_POSITIONS crystal_sg Fe 0.362400000 0.279900000 0.250000000 P 0.587500000 0.798900000 0.250000000 Na 0.740900000 0.251300000 0.002200000 Na 0.917800000 0.738500000 0.250000000 C 0.060500000 0.234500000 0.250000000 O 0.123300000 0.461600000 0.250000000 O 0.143000000 0.031700000 0.250000000 O 0.919500000 0.215700000 0.250000000 O 0.321000000 0.281700000 0.565500000 O 0.431600000 0.674000000 0.250000000 O 0.569400000 0.096500000 0.250000000 K_POINTS automatic 6 12 10 1 1 1
Lammps
To use lammps you will need to load the lammps module
module load lammps
and use a submission file similar to the following
- lammps.sub
#SBATCH -J Zn-ZnO-rapid #SBATCH --ntasks=4 #SBATCH --mail-type=FAIL #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --time=02:00:00 #SBATCH --export=ALL module load lammps/23Oct2017 mpirun lmp_forge < in.heating
Other options may be configured. please refer to the lammps documentation .
Version 23Oct2017 has support for the following packages
asphere body class2 colloid compress coreshell dipole granular kspace manybody mc meam misc molecule mpiio opt peri python qeq replica rigid shock snap srd user-cgdna user-cgsdk user-diffraction user-dpd user-drude user-eff user-fep user-lb user-manifold user-meamc user-meso user-mgpt user-misc user-molfile user-netcdf user-omp user-phonon user-qtb user-reaxc user-smtbq user-sph user-tally user-uef
Maple
Default version = 2015
Example:
This is a simple polynomial and the goal is to find the roots of the polynomial.
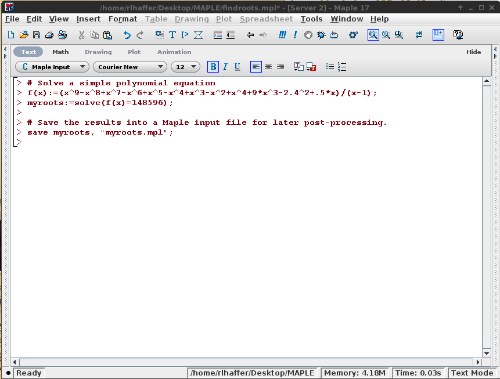
Open Maple on your Windows or Linux workstation and start a new project:
- findroots.mpl
#Solve a simple polynomial equation\\ f(x):=(x^9-x^8+x^7-x^6+x^5-x^4+x^3-x^2+x^4+9*x^3-2.4^2+.5*x)/(x-1);\\ myroots:=solve(f(x)=148596);\\ #Save the results into a Maple input file for later post-processing. save myroots, "myroots.mpl";
When this runs, it will save the output to the myroots.mpl file.
Now, you need to SSH into the Forge with your MST userID/password.
Once connected, you need to copy the findroots.mpl file from your local system to your home directory on the cluster.
You can build your folder structure on the Forge how you wish, just use something that is meaningful.
Once copied, you can return to your terminal on Forge and create your jobfile to run your findroots.mpl project.
Open vi to create your job file.
Since this example is simple, it is assigned 1 node, with 2 processors and 15 minutes wall time.
The job will email when it begins, ends or fails.
First:
module load maple/2015
Then write your jobfile:
- maple.sub
##Example Maple script #!/bin/bash #SBATCH --job-name=maple_example #SBATCH --nodes=1 #SBATCH --ntasks=2 #SBATCH --mem=2000 #SBATCH --time=00:15:00 #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-type=FAIL #SBATCH --mail-user=rlhaffer@mst.edu maple -q findroots.mpl
Then run:
sbatch maple.sbatch
When the job finishes, you will have a
slurm-jobID.out
This contains the output of the finished job.
Matlab
Matlab is available to run in batch form or interactively on the cluster, we currently have versions 2012, 2013a, and 2014a installed through modules.
Interactive Matlab
The simplest way to get up and running with matlab on the cluster is to simply run
matlab
from the login node. This will start an interactive job on the backend nodes, load the 2014a module and open matlab. If you have connected with X forwarding you will get the full matlab GUI to use however you would like. This method however limits you to 1 core for 4 hours maximum on one of our compute nodes. To use more than 1 core, or run for longer than 4 hours, you will need to submit your job as a batch submission and use Matlab's Distributive Computing Engine.
Batch Submit
If you want to use Batch Submissions for Matlab you will need to create a submission script similar to the ones above in quick start, but you will want to limit the nodes your job runs on 1, please see the sample submission script below.
- matlab.sub
#!/bin/bash #SBATCH --nodes=1 #SBATCH --ntasks=12 #SBATCH -J Matlab_job #SBATCH -o Forge-%j.out #SBATCH --time=01:00:00 module load matlab matlab < helloworld.m
This submission asks for 12 processors on 1 node for an hour, the maximum per node we currently have is 64. If you load the any version older than 2014a your matlab can't use more than 12 processors as that is what mathworks limited you to using on one machine, however Mathworks lifted this limitation to 512 processors in 2014a.
Distributive Computing Engine
Mathworks has developed a parallel computing product that the campus has purchased licenses for, We have developed a parallel cluster profile that must be loaded on a user to user basis. To begin using the distributive computing profile you must open an interactive matlab session with X forwarding so you get the matlab GUI.
[blspcy@login-44-0 ~]$ matlab
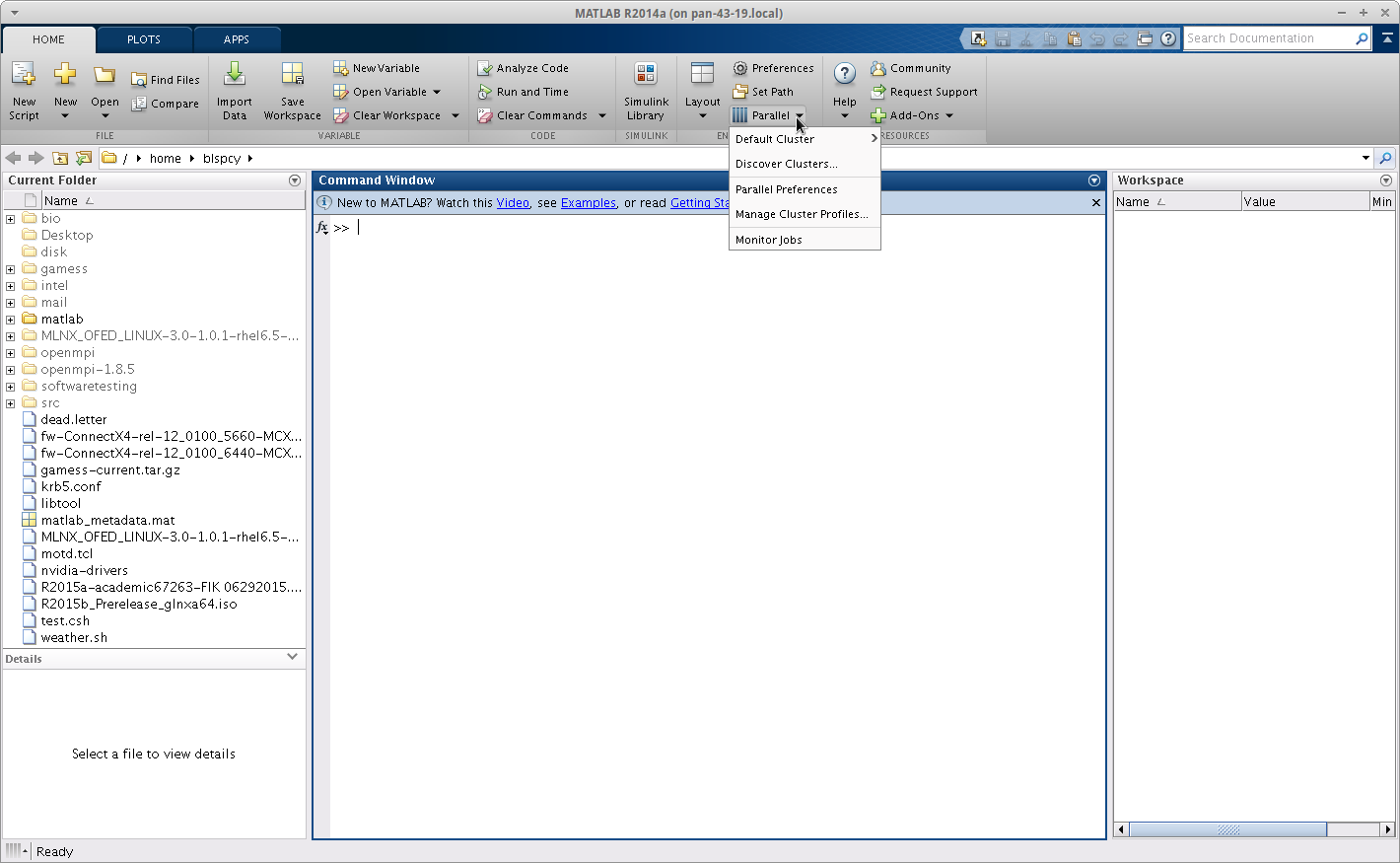
Once the GUI is open you need to open the parallel computing options menu, noted simply by parallel.
Now open the Parallel profile manager and import the Forge.settings profile from the root directory of the version of matlab you are using, e.g. for 2014a /share/apps/matlab/matlab-2014/
You will now have a Forge_Import profile in your profile list, select this profile and set it as default.
If you would like you may validate the imported profile and see that it is indeed working.
The validation will run a series of five tests which it should pass, if for any reason the profile doesn't pass any test please contact the helpdesk or create a ticket at http://help.mst.edu
To modify how many workers the pool will try to use when you call
matlabpool open
in your matlab script you may edit the profile and set the NumWorkersRange value to the number you wish to use, or simply specify when you call the pool open
matlabpool open 24
will open a pool of 24 workers.
You may now use matlabpool open in your code and it will call the Forge cluster profile configured above, These steps must be followed for every version of Matlab you wish to use before calling the pool. The parallel profiles are only built for versions newer than 2014a.
To make use of this new found power you must implement
matlabpool open
calls in your matlab script then run it using either the interactive method or batch submit method listed above, make sure to close the pool when you are finished with your work with
matlabpool close
Keep in mind that the University only has 160 of these licenses and your job may fail if someone else is using the licenses when your job runs.
METIS
\*\*\Under Construction/*/*/
METIS and parMETIS are installed as modules, to use the software please load the modules with either
module load metis
or
module load parmetis
Once loaded you may submit jobs using the METIS or parMETIS binaries as defined in the METIS user manual or the parMETIS user manual
MSC
Nastran
MSC nastran 2014.1 is available through the module msc/nastran/2014
module load msc/nastran/2014
Here is an example job file.
- msc.sub
#!/bin/bash #SBATCH -J Nastran #SBATCH -o Forge-%j.out #SBATCH --ntasks=1 #SBATCH --time=01:00:00 nast20141 test.bdf #you should also be able to use .dat files with nastran this way.
Submit it as normal.
NAMD
This article is under development. Things referenced below are written as we make progress and some instruction may not work as intended.
Below is an example file for namd, versions prior to 2.12b have not been tested as of yet and may not function in the same way as 2.12b.
- namd.sub
#SBATCH -J NAMD_EXAMPLE #SBATCH --ntask=16 #SBATCH --time=01:00:00 #SBATCH -o Forge-%J.out module load namd/2.12b charmrun +p16 namd2 [options]
You will need to give namd2 whatever options you would like to run, user documentation for what those may be is available from the developers at http://www.ks.uiuc.edu/Research/namd/2.12b1/ug/.
Gamess
To use gamess you will need to load the gamess module
module load gamess
and use a submission file similar to the following
- gamess.sub
#!/bin/bash #SBATCH -J Gamess_test #SBATCH --time=01:00:00 #SBATCH --ntasks=2 #SBATCH -o Forge-%J.out rungms -i ./test.inp
We have customized the rungms script extensively for our system and have introduced several useful switch options.
| Switch | Function |
| -i | defines the path to the input file, no default value |
| -d | defines the path to the data folder, default value is the submission directory |
| -s | defines the path to the scratch folder, default value is ~/gamess/scr-$input-$date this folder gets removed once the job completes. |
| -b | defines the binary that will be used, default is /share/apps/gamess/gamess.00.x useful if you've compiled your own version of gamess. |
| -a | defines the auxdata folder, default is /share/apps/gamess/auxdata useful if you've compiled your own version of gamess. |
OpenFOAM
First load the module
module load OpenFOAM
then source the bash file,
source $FOAMBASH
or the csh file if you are using csh as your shell
source $FOAMCSH
Then you may use open foam to work on your data files, anything that needs to be generated interactively needs to be done in an interactive job
sinteractive
You can create a job file to submit from the data directory to do anything that needs more processing, please see the example below.
- foam.sub
#!/bin/bash #SBATCH -J FOAMtest #SBATHC --nodes=1 #SBATCH --ntasks=2 #SBATCH --time=01:00:00 blockMesh
This will execute blockMesh on the input files in the submission directory on 1 node with 2 CPUs for 1 hour.
Python
Python 2
Python modules are available through system modules as well. To see what python modules have been installed issue
module avail python
These are the site installed python modules and can be used just like any other module.
module load python
will load all python modules,
module load python/matplotlib
will load the matplotlib module and any module it depends on.
Python 3
There are many modules available for Python 3. However, unlike Python 2.7, all of the python modules are installed in the python directory rather than being separate modules. Currently, the newest Python 3 version available on the Forge is Python 3.6.4. To see a current list of available python versions, run the command
module avail python
To see a list of all available Python modules available for a particular Python version, load that Python version and run
pip list
Users are welcome to install additional modules on their account using
pip install --user <package name>
QMCPack
Start by loading the module.
module load qmcpack
Then create your submission file in the directory where your input files are, see example below.
- qmcpack.sub
#!/bin/bash #SBATCH -J qmcpack #SBATCH -o Forge-%j.out #SBATCH --ntasks=8 #SBATCH --time=1:00:00 #SBATCH --constraint=intel mpirun qmcapp input.xml
Note: QMCPack is compiled to run only on intel processors, you need to add
--constraint=intel
to make sure that your job doesn't run on a node with amd processors.
This should run qmcpack on 8 cpus for 1 hour on your input file. If you want to run multiple instances, in parallel, in the same job you may pass an input.list file in place of the input.xml which contains all the input xml files you wish to run.
For more information on creating the input files please see QMCPack's user guide
SAS 9.4
Notes:
SAS can be run from the login node, with an X session, or from command line.
SAS is node-locked to only forge.mst.edu, so it won't work in a jobfile. You must run it interactively on forge.mst.edu.
For both methods, you need to load the SAS module.
module load SAS/9.4
Interactive:
Steps:
- Start a local X Server XWin 32
- SSH to forge.mst.edu
- On our Linux builds, type the command “ssh -XC forge.mst.edu”
- Once logged in onto the Forge, run
sinteractive -p=free
- the sinteractive command will open a new SSH tunnel on a compute node with free resources.
- One this new SSH command line, run the
sas_en
and it will open the interface on the compute node that sinteractive has set.
- Remember to close all your windows when you're finished
Jobfile
- sas.sub
#!/bin/bash #SBATCH --job-name=job_name #SBATCH --ntasks=1 #SBATCH --time=23:59:59 #SBATCH --mem=31000 #SBATCH --mail-type=begin,end,fail,requeue #SBATCH --export=all #SBATCH --out=Forge-%j.out module load SAS/9.4 sas_en -nodms sas_script.sas
Command line:
sas_en -nodms
runs the command line version of sas. (type endsas; to get out)
[rlhaffer@login-44-0 apps]$ sas_en -nodms
NOTE: Copyright (c) 2002-2012 by SAS Institute Inc., Cary, NC, USA.
NOTE: SAS (r) Proprietary Software 9.4 (TS1M3)
Licensed to THE CURATORS OF THE UNIV OF MISSOURI - T&R, Site 70084282.
NOTE: This session is executing on the Linux 2.6.32-431.11.2.el6.x86_64 (LIN
X64) platform.
SAS/QC 14.1
NOTE: Additional host information:
Linux LIN X64 2.6.32-431.11.2.el6.x86_64 #1 SMP Tue Mar 25 19:59:55 UTC 2014
x86_64 CentOS release 6.5 (Final)
You are running SAS 9. Some SAS 8 files will be automatically converted
by the V9 engine; others are incompatible. Please see
http://support.sas.com/rnd/migration/planning/platform/64bit.html
PROC MIGRATE will preserve current SAS file attributes and is
recommended for converting all your SAS libraries from any
SAS 8 release to SAS 9. For details and examples, please see
http://support.sas.com/rnd/migration/index.html
This message is contained in the SAS news file, and is presented upon
initialization. Edit the file "news" in the "misc/base" directory to
display site-specific news and information in the program log.
The command line option "-nonews" will prevent this display.
NOTE: SAS initialization used:
real time 0.11 seconds
cpu time 0.04 seconds
1?
PRINTING:
- Select File | Print.
- This will create a postscript file, “sasprt.ps”, in the directory you started SAS from. You can download it to your computer and use ps2pdf to print it.
StarCCM+
Engineering Simulation Software
Default version = 10.04.011
Other working versions:
11.02.009
module load starccm/11.02
10.06.010
module load starccm/10.06
8.04.010
module load starccm/8.04
7.04.011
module load starccm/7.04
6.04.016
module load starccm/6.04
Job Submission Information
Copy your .sim file from the workstation to your cluster home profile. The Forge documentation explains how to do this.
Once copied, create your job file.
Example job file: First
module load starccm/10.04
- starccm.sub
#!/bin/bash #SBATCH --job-name=starccm_example #SBATCH --nodes=1 #SBATCH --ntasks=12 #SBATCH --partition=requeue #SBATCH --time=08:00:00 #SBATCH --mail-type=FAIL #SBATCH --mail-type=BEGIN #SBATCH --mail-type=END #SBATCH --mail-user=joeminer@mst.edu module load starccm/10.04 time starccm+ -power -batch -np 12 -licpath 1999@flex.cd-adapco.com -podkey AABBCCDDee1122334455 /path/to/your/starccm/simulation/example.sim
It's prefered that you keep the ntasks and -np set to the same processor count.
Breakdown of the script:
This job will use 1 node, asking for 12 processors, for a total wall time of 8 hours and will email you when the job starts, finishes or fails.
The StarCCM commands:
| -power | using the power session and power on demand key |
| -batch | tells Star to utilize more than one processor |
| -np | number of processors to allocate |
| -licpath | 1999@flex.cd-adapco.com - Since users of StarCCM at S&T must have Power On Demand keys, we use CD-Adapco's license server |
| -podkey | Enter your PoD key….AABBCCDDee1122334455 This is given to you when you sign up for Star, or someone from the FSAE team provides it. |
| /path/to/your/starccm/simulation/example.sim | use the true path to your .sim file in your cluster home directory |
Vasp
To use our site installation of Vasp you must first prove that you have a license to use it by emailing your vasp license confirmation to it-research-support@mst.edu.
Once you have been granted access to using vasp you may load the vasp module
module load vasp
and create a vasp job file, in the directory that your input files are, that will look similar to the one below.
- vasp.sub
#!/bin/bash #SBATCH -J Vasp #SBATCH -o Forge-%j.out #SBATCH --time=1:00:00 #SBATCH --ntasks=8 mpirun vasp
This example will run the standard vasp compilation on 8 cpus for 1 hour.
If you need the gamma only version of vasp use
mpirun vasp_gam
in your submission file.
If you need the non-colinear version of vasp use
mpirun vasp_ncl
in your submission file.
There are some globally available Psudopoetentials available, the module sets the environment variable $POTENDIR to the global directory.